|
Here the default downsampling function mean renders a faithful size-reduced version of the original, with all the raster grid points that overlap a given pixel being averaged to create the final pixel value. The coords argument is optional, but without it the default integer indexing from Numpy would be used, which would not match how this data was generated (sampling over each of the ys).ĭataArrays like da happen to be the format used for datashader’s own rasterized output, so you can now immediately turn this hand-constructed array into an image using tf.shade() just as you would for points or lines rasterized by datashader: Here we are declaring that the first dimension of the DataArray (the rows) is called y and corresponds to the indicated continuous coordinate values in the list ys, and similarly for the second dimension (the columns) called x. 
DataArray ( z, coords = ) da = sample ( f ) meshgrid ( xs, ys ) z = fn ( x, y ) return xr. linspace ( range_, range_, n ) x, y = np. Import numpy as np, datashader as ds, xarray as xr from datashader import transfer_functions as tf, reductions as rd def f ( x, y ): return np. Re-rasterization #įirst, let’s explore the regularly gridded case, declaring a small raster using Numpy and wrapping it up as an xarray DataArray for us to re-rasterize: Fully arbitrary unstructured grids ( Trimeshes) are discussed separately. Datashader also provides fast methods for rasterizing these more general rectilinear or curvilinear grids, known as quadmeshes as described later below. In other cases, your data is stored in a 2D array similar to a raster, but represents values that are not regularly sampled in the underlying coordinate space. Rasterizing into a common grid can help you implement complex cross-datatype analyses or visualizations. Datashader provides fast methods for “regridding”/ ”re-sampling”/”re-rasterizing” your regularly gridded datasets, generating new rasters on demand that can be used together with those it generates for any other data types. Even so, the rasters you have already are not always the ones you need for a given purpose, having the wrong shape, range, or size to be suitable for overlaying with or comparing against other data, maps, and so on. In some cases, your data is already rasterized, such as data from imaging experiments, simulations on a regular grid, or other regularly sampled processes. Z = np.Datashader renders data into regularly sampled arrays, a process known as rasterization, and then optionally converts that array into a viewable image (with one pixel per element in that array). # I'm fairly sure there's a more efficient way of doing this. But if it is, you can avoid having to interpolate: def method_3(): In general, with sampled data, this will not be true. For every possible unique (x,y) pair, there is a corresponding (x,y,z) in your data.įrom this, it follows that the number of (x,y,z) pairs must be equal to the square of the number of unique x points (where the number of unique x positions equals the number of unique y positions).The number of different x sample positions equals the number of different y sample positions.There's a third option, depending on how your (x,y,z) is set up. Method 3: No Interpolation (constraints on sampled data) This method produces the following graphs: Z = iddata((mat, mat), mat, (X,Y), method='nearest') # Depending on your "error", you may be able to use other methods # Interpolate (x,y,z) points over a normal (x,y) grid 
# This works if you have (x,y,z) tuples that you're *not* generating, and (x,y) points Method 2: Interpolating given z points over a regular grid # Method 2: Here, the returned matrix looks like: mat = , Y_err = (np.random.rand(*y.shape) - 0.5) * xy_max_error X_err = (np.random.rand(*x.shape) - 0.5) * xy_max_error # Half of this will be in the + direction, half will be in the - dir. # First we generate the (x,y,z) tuples to imitate "real" data First, I define a function that generates fake data: def gen_fake_data(): If you don't have that ability, and are given a fixed (x,y,z). This is relatively easy, since you can generate z at whatever points you want. # This works if you are generating z, given (x,y) # dim_? is the granularity in that direction If you just have just a list of ( x, y, z) tuples, it's harder (see method_2() below, and maybe method_3()).Ĭonstants # min_? is minimum bound, max_? is maximum bound, you know the formula for it) it's very easy (see method_1() below). 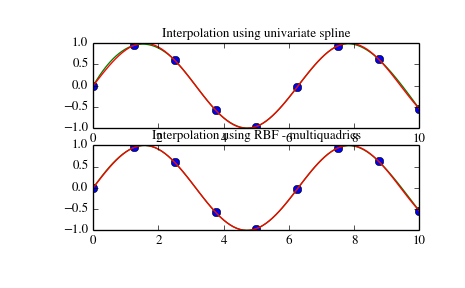
Depending on whether you're generating z or not, you have at least two different options.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
